import pandas as pd
import geopandas as gpd
from sklearn.cluster import KMeans
from sklearn.preprocessing import StandardScaler
# Example dataset (San Diego tracts)
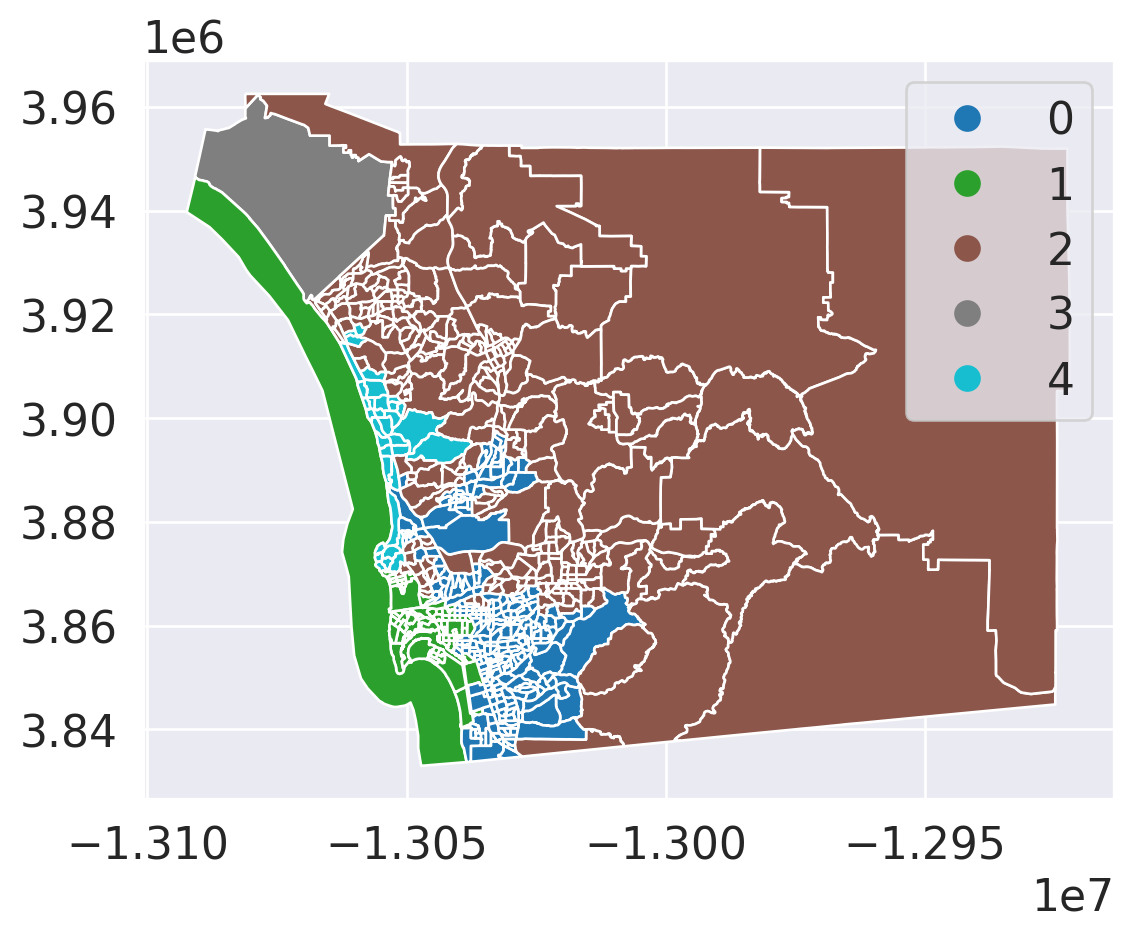
data = gpd.read_file('~/data/385/sandiego_tracts.gpkg')
# Select clustering variables
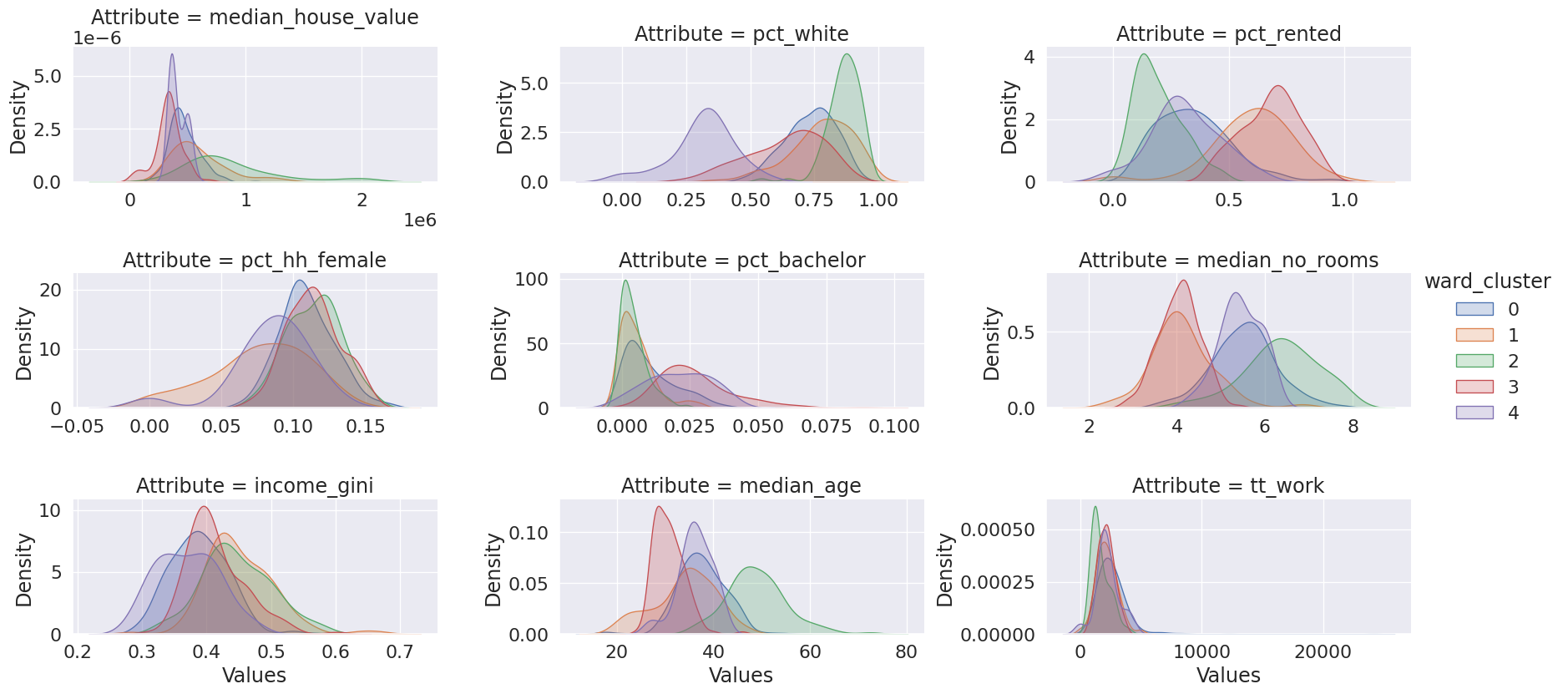
cluster_variables = ["median_house_value", "pct_white", "pct_rented", "pct_hh_female", "pct_bachelor", "median_no_rooms", "income_gini", "median_age", "tt_work"]
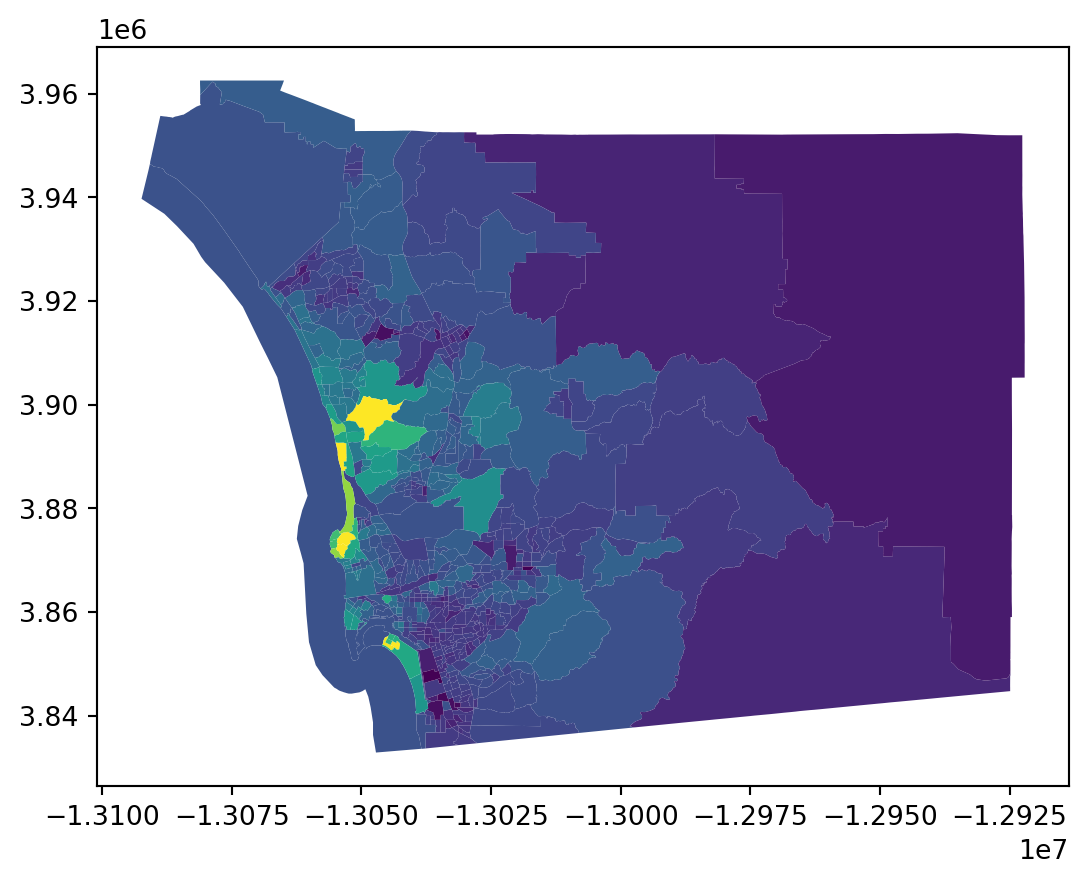
data.plot('median_house_value')