Spatial Weights
Spatial Weights
Spatial weights are mathematical structures used to represent spatial relationships. Many spatial analytics, such as spatial autocorrelation statistics and regionalization algorithms rely on spatial weights. Generally speaking, a spatial weight \(w_{i,j}\) expresses the notion of a geographical relationship between locations \(i\) and \(j\). These relationships can be based on a number of criteria including contiguity, geospatial distance and general distances.
libpysal offers functionality for the construction, manipulation, analysis, and conversion of a wide array of spatial weights.
We begin with construction of weights from common spatial data formats.
Imports
There are functions to construct weights directly from a file path.
Weight Types
- Contiguity
- Distance Based Weights
Contiguity Weights
Contiguity Weights
- Alternative Definitions
- Queen
- Rook
- Bishop
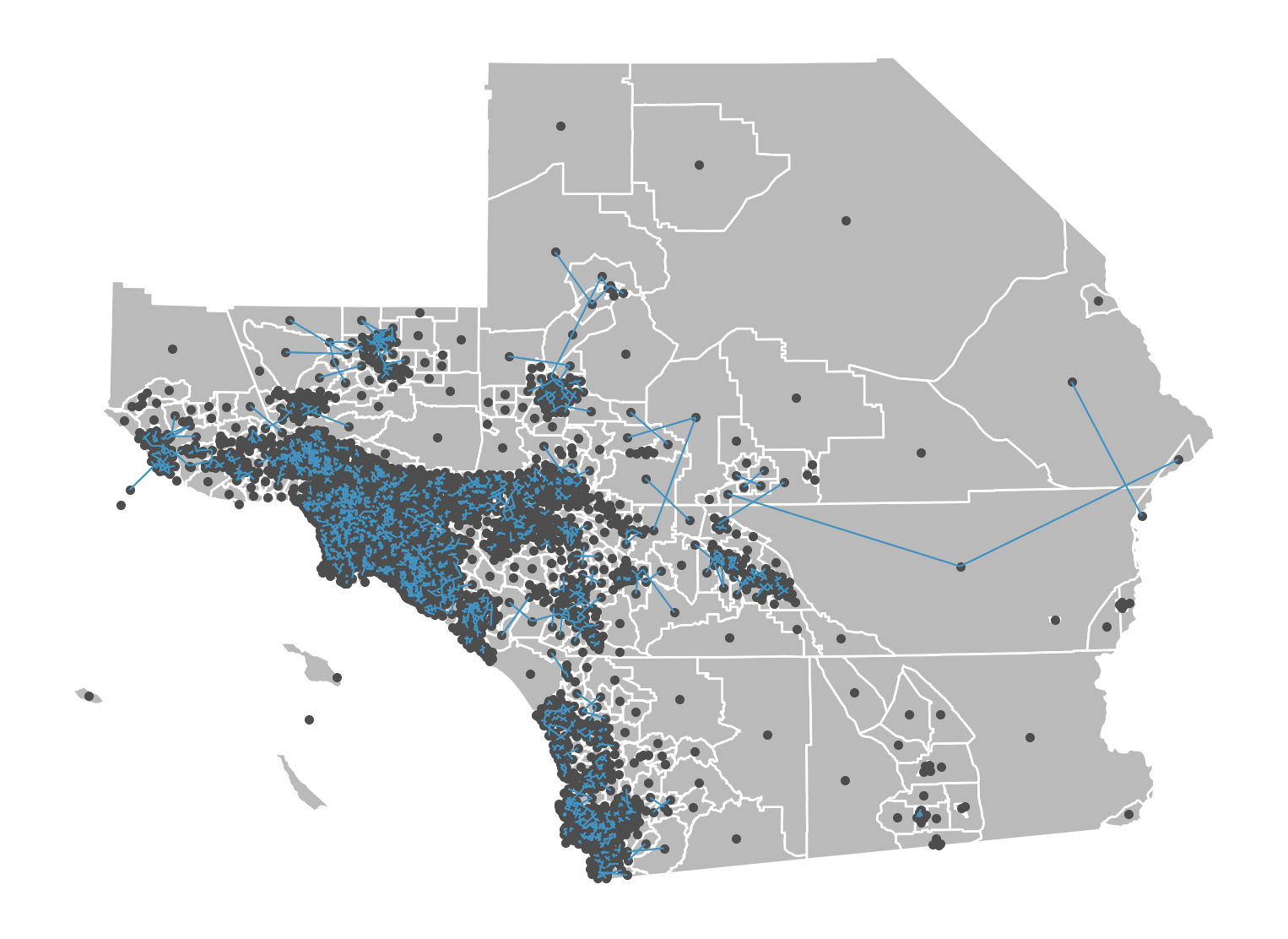
Queen Weights
A commonly-used type of weight is a queen contigutiy weight, which reflects adjacency relationships as a binary indicator variable denoting whether or not a polygon shares an edge or a vertex with another polygon. These weights are symmetric, in that when polygon \(A\) neighbors polygon \(B\), both \(w_{AB} = 1\) and \(w_{BA} = 1\).
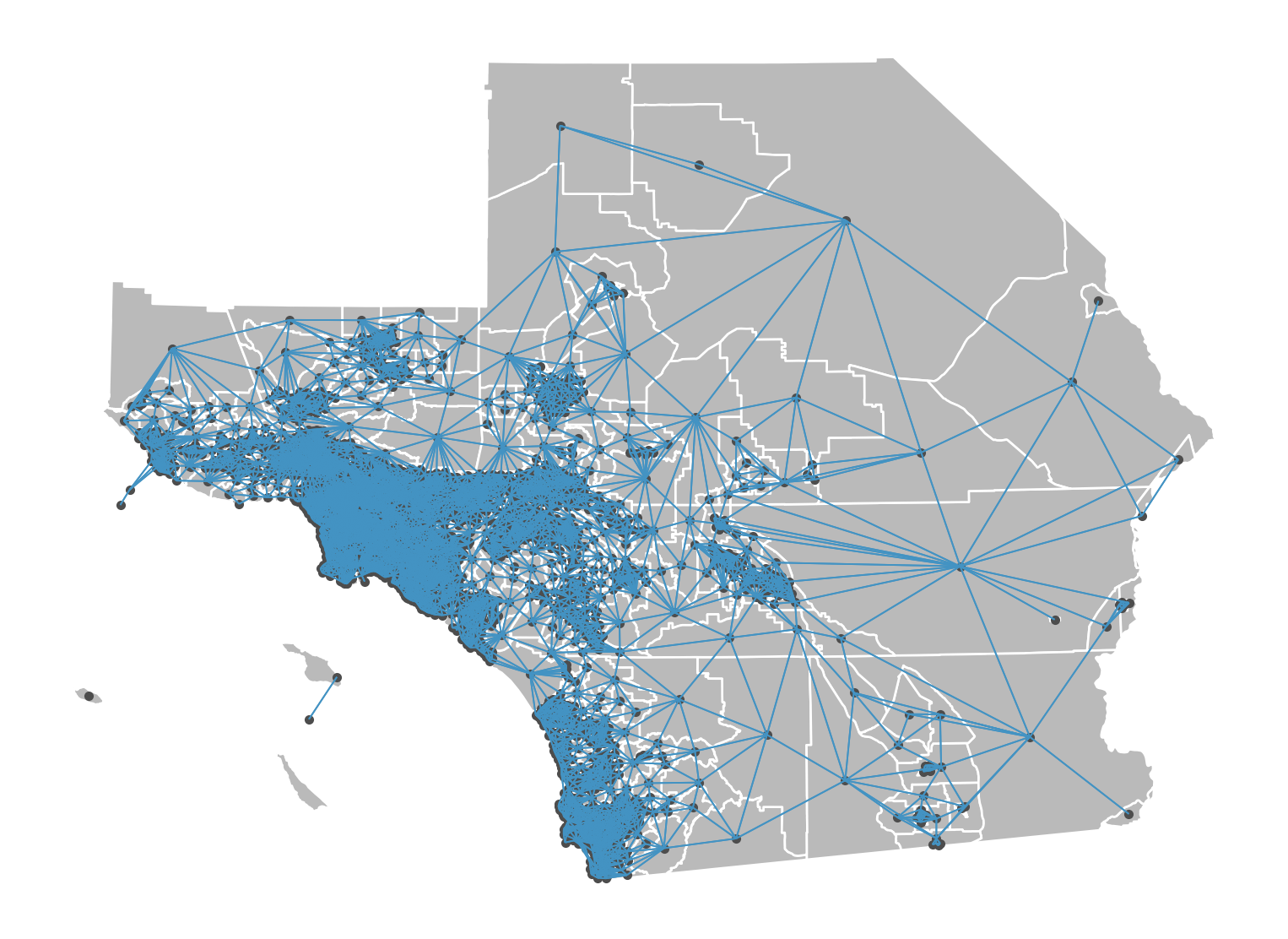
Constructing Queen Weights
To construct queen weights from a shapefile, we will use geopandas to read the file into a GeoDataFrame, and then use libpysal to construct the weights:
Weight Attributes
All weights objects have a few traits that you can use to work with the weights object, as well as to get information about the weights object.
To get the neighbors & weights around an observation, use the observation’s index on the weights object, like a dictionary:
Weights Matrix
A full, dense matrix describing all of the pairwise relationships is constructed using the .full method, or when libpysal.weights.full is called on a weights object:
Number of Neighbors
Note that this matrix is binary, in that its elements are either zero or one, since an observation is either a neighbor or it is not a neighbor.
Standardization
However, many common use cases of spatial weights require that the matrix is row-standardized. This is done simply in PySAL using the .transform attribute
Now, if we build a new full matrix, its rows should sum to one:
Sparse Weights
Since weight matrices are typically very sparse, there is also a sparse weights matrix constructor:
<Compressed Sparse Row sparse matrix of dtype 'float64'
with 29374 stored elements and shape (4580, 4580)>Neighbors
Let’s look at the neighborhoods of the 101th observation
Plotting
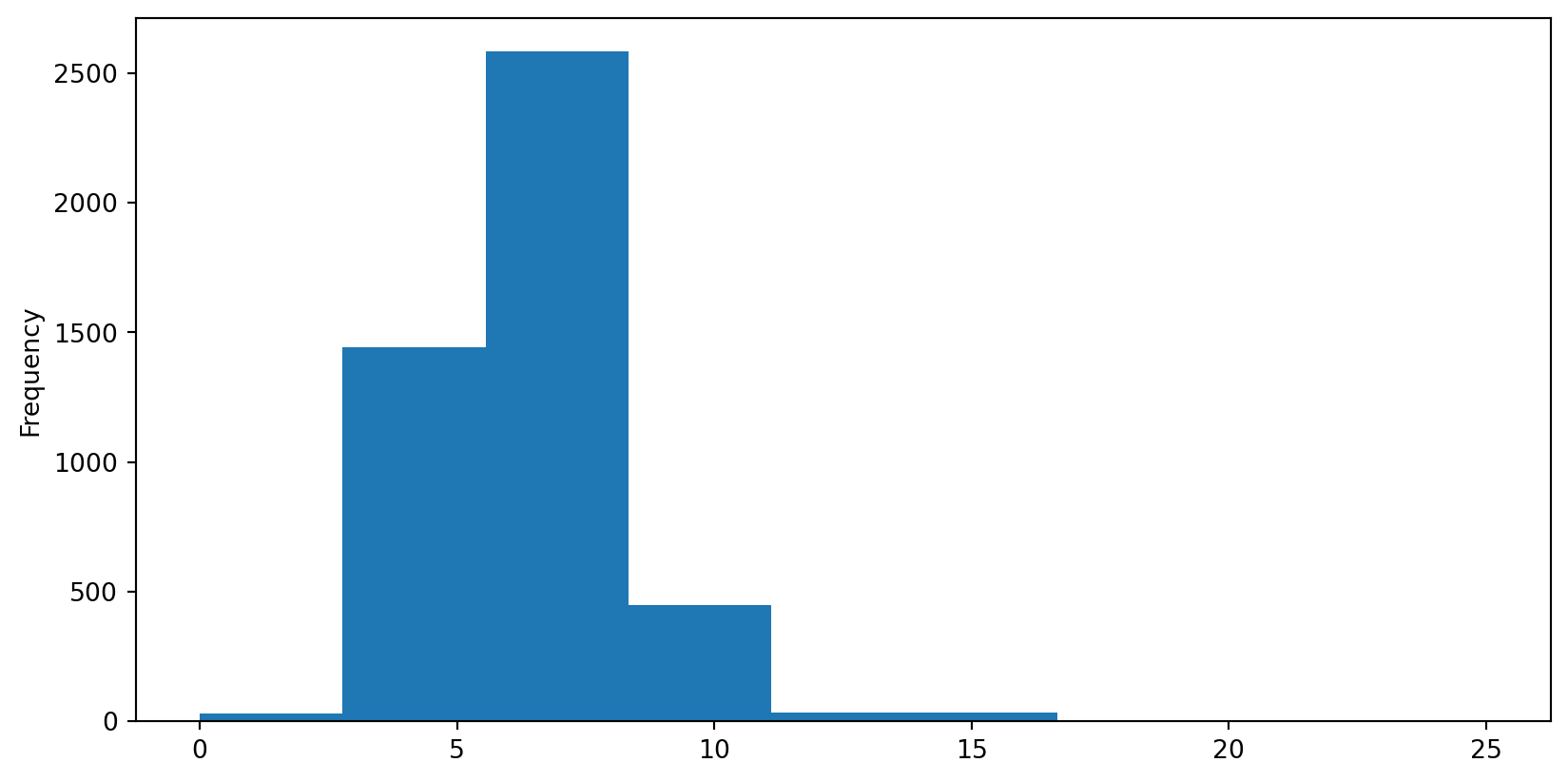
Connectivity Histogram
Cardinalities
dict_values([9, 9, 4, 7, 7, 5, 5, 6, 5, 8, 9, 8, 4, 3, 5, 5, 6, 6, 4, 5, 5, 6, 7, 9, 6, 4, 7, 8, 7, 5, 7, 2, 6, 6, 8, 3, 7, 7, 5, 8, 6, 5, 5, 4, 6, 6, 7, 7, 4, 6, 7, 4, 5, 6, 13, 6, 7, 6, 8, 6, 6, 6, 2, 6, 6, 8, 6, 7, 7, 6, 3, 5, 6, 6, 3, 7, 6, 5, 5, 5, 8, 8, 6, 8, 9, 7, 7, 6, 7, 5, 5, 7, 7, 6, 5, 7, 8, 8, 4, 7, 5, 4, 4, 6, 9, 6, 6, 7, 4, 8, 6, 6, 5, 6, 6, 6, 6, 7, 6, 8, 6, 6, 6, 8, 6, 6, 5, 8, 4, 5, 7, 5, 5, 5, 5, 5, 2, 4, 4, 7, 6, 8, 6, 9, 4, 6, 7, 5, 5, 6, 6, 4, 8, 9, 7, 4, 8, 4, 6, 5, 5, 4, 5, 6, 7, 8, 4, 6, 5, 6, 6, 6, 4, 6, 7, 6, 5, 6, 7, 6, 7, 7, 7, 7, 3, 10, 6, 6, 7, 7, 5, 6, 7, 8, 6, 5, 9, 7, 9, 6, 6, 4, 6, 6, 5, 7, 7, 7, 6, 4, 7, 8, 7, 5, 6, 6, 4, 6, 6, 5, 9, 7, 5, 7, 4, 7, 7, 3, 6, 7, 5, 5, 6, 6, 5, 4, 6, 5, 5, 6, 5, 10, 4, 3, 6, 1, 8, 6, 4, 5, 5, 7, 6, 4, 7, 4, 5, 6, 6, 5, 10, 3, 5, 5, 9, 5, 7, 5, 5, 7, 5, 8, 4, 6, 5, 7, 7, 7, 5, 6, 7, 5, 3, 7, 5, 4, 6, 3, 5, 6, 5, 5, 5, 4, 4, 7, 7, 5, 5, 5, 7, 9, 6, 4, 4, 5, 7, 4, 4, 7, 4, 6, 6, 4, 8, 6, 7, 5, 8, 6, 7, 6, 8, 8, 4, 5, 7, 6, 3, 5, 5, 4, 6, 6, 7, 5, 5, 5, 3, 5, 7, 6, 8, 5, 5, 5, 7, 6, 6, 7, 3, 4, 8, 4, 7, 4, 6, 6, 4, 4, 4, 5, 3, 5, 6, 4, 6, 7, 8, 5, 6, 6, 6, 7, 5, 8, 5, 3, 5, 5, 6, 4, 7, 7, 7, 4, 6, 5, 6, 9, 4, 7, 5, 5, 5, 7, 4, 7, 8, 7, 7, 6, 5, 5, 7, 5, 9, 5, 5, 6, 4, 8, 6, 4, 5, 5, 5, 7, 6, 6, 3, 6, 7, 5, 4, 5, 5, 7, 7, 5, 4, 6, 6, 6, 5, 5, 5, 17, 7, 6, 6, 14, 5, 5, 6, 6, 4, 11, 7, 6, 4, 6, 5, 7, 6, 6, 4, 7, 4, 6, 4, 7, 7, 6, 8, 5, 6, 8, 6, 5, 8, 8, 6, 4, 5, 5, 3, 6, 4, 6, 6, 6, 5, 6, 4, 4, 6, 7, 7, 8, 4, 5, 17, 7, 5, 6, 5, 7, 8, 6, 4, 7, 5, 7, 6, 4, 4, 5, 5, 4, 4, 4, 5, 3, 5, 4, 5, 3, 4, 5, 7, 7, 4, 7, 5, 4, 5, 12, 9, 9, 5, 4, 6, 7, 6, 6, 3, 8, 4, 8, 4, 6, 8, 4, 6, 5, 9, 6, 7, 4, 4, 8, 7, 4, 8, 5, 5, 7, 6, 5, 7, 4, 8, 6, 5, 8, 6, 6, 6, 6, 4, 5, 6, 8, 5, 8, 5, 8, 5, 8, 8, 6, 8, 7, 6, 7, 6, 5, 5, 7, 9, 3, 6, 8, 8, 7, 8, 5, 4, 6, 5, 6, 9, 7, 6, 7, 8, 3, 6, 6, 4, 7, 6, 5, 3, 5, 5, 8, 6, 8, 3, 8, 6, 8, 6, 6, 5, 8, 4, 8, 8, 7, 4, 5, 6, 6, 7, 7, 7, 5, 7, 4, 8, 6, 8, 8, 6, 4, 5, 7, 6, 6, 7, 5, 7, 8, 4, 6, 7, 6, 6, 8, 7, 7, 4, 4, 8, 7, 7, 8, 11, 6, 7, 6, 10, 5, 6, 6, 6, 4, 5, 5, 8, 7, 4, 7, 6, 8, 6, 7, 6, 7, 7, 6, 6, 5, 6, 6, 6, 5, 7, 9, 4, 5, 6, 6, 5, 6, 7, 5, 10, 5, 7, 4, 5, 4, 5, 5, 5, 6, 10, 6, 6, 6, 5, 6, 7, 4, 4, 6, 6, 8, 6, 7, 8, 8, 6, 3, 5, 6, 5, 7, 5, 5, 7, 8, 6, 7, 5, 5, 6, 6, 6, 5, 6, 6, 8, 5, 8, 4, 6, 8, 4, 5, 3, 5, 5, 8, 5, 8, 7, 8, 7, 6, 10, 5, 7, 4, 4, 3, 9, 4, 6, 5, 4, 7, 7, 6, 9, 4, 6, 5, 5, 7, 5, 5, 9, 5, 6, 6, 5, 6, 5, 7, 5, 8, 8, 6, 7, 5, 7, 5, 5, 7, 7, 4, 7, 8, 6, 8, 7, 5, 7, 10, 5, 4, 7, 5, 6, 7, 7, 5, 5, 4, 6, 5, 6, 6, 7, 5, 7, 7, 6, 6, 5, 6, 4, 4, 6, 5, 6, 5, 6, 6, 5, 4, 5, 9, 9, 4, 5, 8, 6, 8, 5, 7, 7, 6, 6, 4, 9, 8, 6, 10, 5, 7, 6, 3, 6, 6, 8, 7, 5, 7, 5, 6, 8, 5, 6, 5, 5, 7, 4, 5, 4, 4, 5, 4, 5, 7, 5, 7, 6, 5, 7, 7, 6, 7, 6, 7, 7, 6, 7, 4, 4, 6, 7, 4, 6, 7, 8, 6, 8, 8, 2, 7, 4, 9, 5, 7, 7, 4, 6, 6, 4, 5, 5, 5, 9, 6, 8, 6, 9, 9, 7, 7, 5, 7, 5, 5, 5, 5, 4, 7, 6, 5, 6, 9, 6, 4, 6, 5, 6, 7, 6, 6, 4, 5, 3, 7, 4, 5, 6, 10, 4, 7, 6, 4, 6, 6, 8, 5, 6, 6, 7, 6, 6, 4, 7, 6, 7, 8, 6, 6, 7, 9, 6, 6, 9, 6, 8, 3, 5, 7, 8, 9, 7, 5, 5, 7, 7, 7, 6, 5, 6, 7, 7, 3, 6, 6, 5, 9, 5, 8, 7, 5, 6, 5, 7, 7, 7, 6, 6, 8, 6, 5, 6, 7, 3, 6, 6, 8, 7, 6, 8, 5, 7, 6, 5, 6, 7, 4, 4, 7, 5, 20, 6, 5, 6, 6, 6, 7, 5, 6, 7, 5, 4, 5, 4, 9, 4, 7, 6, 8, 7, 5, 5, 5, 5, 7, 5, 6, 6, 5, 3, 5, 4, 6, 8, 6, 2, 7, 5, 5, 5, 6, 8, 7, 7, 7, 8, 7, 6, 5, 5, 6, 5, 6, 5, 7, 6, 7, 4, 6, 4, 5, 7, 5, 9, 6, 7, 7, 5, 10, 8, 5, 5, 5, 7, 6, 6, 7, 6, 7, 6, 5, 4, 5, 5, 6, 7, 10, 4, 4, 6, 6, 8, 7, 5, 6, 6, 9, 5, 5, 6, 6, 8, 5, 7, 6, 4, 9, 7, 5, 5, 6, 5, 16, 5, 4, 6, 4, 8, 8, 6, 6, 7, 7, 5, 10, 7, 6, 7, 6, 7, 5, 7, 5, 7, 6, 7, 6, 4, 8, 6, 7, 6, 6, 5, 4, 4, 5, 9, 4, 8, 8, 7, 6, 6, 11, 6, 4, 6, 6, 4, 7, 5, 6, 7, 5, 5, 3, 7, 5, 5, 3, 6, 6, 9, 5, 8, 8, 3, 6, 5, 5, 8, 5, 5, 7, 5, 9, 8, 9, 6, 6, 6, 7, 8, 8, 7, 6, 7, 9, 4, 8, 9, 9, 7, 7, 8, 6, 5, 6, 5, 5, 6, 5, 5, 8, 6, 7, 5, 7, 4, 7, 5, 10, 7, 8, 5, 6, 9, 9, 9, 10, 4, 5, 6, 9, 4, 5, 6, 6, 5, 6, 8, 9, 9, 9, 10, 7, 9, 5, 3, 6, 7, 7, 9, 7, 8, 9, 8, 7, 8, 6, 7, 6, 9, 9, 5, 8, 3, 3, 7, 5, 7, 9, 9, 8, 7, 7, 10, 6, 8, 7, 6, 8, 6, 8, 5, 5, 5, 4, 5, 7, 8, 9, 7, 3, 6, 4, 7, 7, 7, 6, 10, 6, 7, 8, 7, 10, 9, 7, 6, 8, 5, 7, 7, 6, 5, 5, 5, 7, 5, 6, 4, 5, 6, 4, 9, 7, 4, 6, 4, 9, 9, 6, 8, 5, 9, 6, 9, 6, 7, 7, 6, 5, 10, 10, 7, 7, 7, 7, 8, 6, 3, 7, 4, 5, 6, 4, 10, 9, 5, 8, 6, 9, 5, 7, 5, 7, 5, 8, 8, 9, 9, 13, 7, 7, 8, 6, 1, 7, 6, 5, 6, 7, 9, 5, 7, 6, 4, 4, 3, 7, 8, 6, 7, 8, 6, 4, 6, 9, 8, 6, 6, 6, 8, 7, 7, 5, 5, 2, 3, 7, 6, 6, 6, 8, 5, 5, 6, 4, 5, 7, 8, 6, 9, 4, 5, 5, 3, 5, 10, 5, 15, 4, 6, 9, 8, 8, 7, 5, 8, 7, 8, 9, 8, 7, 5, 6, 8, 4, 9, 6, 9, 6, 6, 6, 5, 7, 10, 11, 11, 6, 5, 6, 7, 4, 5, 11, 16, 5, 7, 9, 7, 6, 7, 7, 6, 5, 9, 6, 10, 7, 5, 7, 6, 5, 3, 8, 3, 11, 5, 5, 9, 9, 7, 6, 6, 8, 10, 6, 10, 4, 8, 7, 4, 7, 4, 5, 9, 4, 5, 9, 9, 6, 6, 8, 9, 9, 4, 7, 10, 16, 7, 11, 7, 11, 5, 8, 9, 5, 11, 7, 5, 7, 5, 5, 7, 8, 9, 5, 11, 11, 7, 8, 6, 5, 6, 5, 7, 9, 4, 7, 8, 7, 8, 4, 6, 7, 9, 9, 8, 6, 4, 7, 4, 6, 7, 6, 8, 6, 7, 4, 8, 5, 7, 7, 10, 7, 7, 6, 4, 6, 6, 13, 3, 8, 9, 6, 4, 6, 5, 6, 12, 5, 2, 6, 5, 7, 6, 7, 8, 5, 7, 4, 6, 7, 10, 3, 17, 10, 8, 6, 5, 7, 14, 1, 12, 8, 4, 4, 8, 8, 4, 8, 6, 8, 6, 5, 3, 8, 6, 7, 8, 6, 9, 8, 5, 11, 5, 3, 7, 5, 4, 5, 3, 7, 7, 6, 5, 8, 2, 5, 6, 11, 8, 6, 6, 7, 6, 7, 6, 6, 4, 4, 6, 9, 6, 9, 6, 5, 4, 3, 5, 8, 7, 10, 4, 6, 7, 4, 6, 6, 4, 6, 4, 8, 6, 10, 8, 5, 8, 6, 4, 6, 6, 5, 11, 4, 7, 8, 7, 6, 5, 5, 10, 16, 8, 5, 7, 9, 3, 7, 5, 3, 4, 8, 6, 6, 7, 6, 9, 7, 7, 6, 6, 11, 7, 7, 8, 6, 6, 7, 4, 7, 6, 3, 7, 7, 6, 8, 15, 7, 6, 4, 6, 9, 6, 6, 6, 10, 7, 8, 4, 9, 11, 6, 6, 6, 4, 11, 6, 5, 5, 5, 5, 7, 5, 5, 5, 4, 4, 8, 4, 6, 5, 7, 8, 6, 7, 5, 6, 7, 7, 6, 4, 10, 5, 6, 7, 5, 4, 7, 5, 6, 5, 7, 5, 5, 10, 6, 5, 5, 3, 7, 5, 6, 6, 6, 7, 5, 4, 7, 6, 11, 6, 8, 6, 9, 5, 14, 4, 5, 13, 8, 6, 7, 4, 7, 5, 9, 5, 8, 4, 7, 7, 8, 7, 4, 7, 5, 10, 8, 9, 5, 7, 6, 4, 7, 8, 8, 12, 7, 6, 8, 10, 5, 7, 6, 6, 6, 5, 5, 5, 6, 9, 5, 7, 4, 7, 8, 5, 7, 6, 5, 7, 7, 4, 4, 6, 7, 7, 6, 8, 6, 5, 4, 6, 5, 5, 4, 6, 6, 6, 7, 5, 4, 6, 3, 6, 7, 1, 7, 11, 4, 4, 12, 7, 6, 8, 5, 7, 8, 3, 6, 5, 5, 5, 9, 6, 9, 6, 5, 3, 7, 6, 8, 8, 9, 7, 5, 7, 6, 6, 5, 5, 10, 4, 8, 5, 8, 4, 5, 6, 5, 14, 8, 1, 5, 6, 7, 8, 10, 4, 7, 5, 9, 5, 5, 7, 4, 4, 7, 5, 8, 6, 9, 6, 8, 7, 4, 6, 4, 7, 5, 4, 6, 3, 7, 6, 6, 7, 11, 8, 6, 7, 8, 5, 7, 7, 9, 6, 5, 6, 6, 8, 6, 9, 7, 7, 7, 8, 5, 5, 5, 9, 6, 5, 4, 7, 7, 5, 7, 5, 5, 6, 8, 6, 5, 6, 4, 6, 6, 7, 7, 5, 6, 9, 6, 5, 6, 5, 8, 9, 7, 11, 7, 11, 15, 6, 7, 8, 3, 10, 8, 10, 8, 8, 4, 6, 5, 7, 6, 5, 6, 6, 7, 5, 9, 9, 9, 7, 11, 6, 6, 6, 5, 6, 6, 8, 6, 7, 5, 7, 8, 10, 10, 9, 7, 3, 9, 10, 7, 7, 6, 10, 6, 6, 7, 4, 7, 4, 5, 6, 6, 5, 5, 7, 8, 7, 4, 7, 7, 7, 7, 7, 7, 7, 6, 7, 6, 7, 7, 8, 7, 7, 5, 8, 10, 8, 7, 7, 6, 6, 8, 7, 6, 5, 6, 6, 9, 6, 2, 7, 7, 7, 7, 6, 7, 7, 4, 9, 12, 5, 5, 5, 5, 4, 9, 5, 7, 6, 7, 6, 4, 8, 7, 7, 7, 3, 5, 6, 5, 5, 7, 8, 6, 9, 4, 5, 6, 7, 5, 4, 6, 9, 5, 6, 10, 10, 6, 8, 5, 6, 4, 7, 8, 8, 5, 4, 9, 6, 4, 6, 5, 7, 5, 7, 6, 4, 10, 5, 6, 5, 5, 4, 6, 4, 6, 16, 8, 3, 7, 6, 5, 5, 5, 7, 7, 6, 9, 7, 6, 6, 6, 8, 3, 8, 4, 5, 5, 6, 6, 8, 9, 4, 7, 8, 7, 5, 6, 6, 6, 12, 4, 5, 6, 7, 7, 4, 8, 5, 7, 6, 7, 7, 9, 8, 7, 6, 7, 7, 6, 7, 5, 6, 3, 6, 5, 3, 6, 10, 7, 6, 6, 6, 6, 13, 10, 6, 8, 4, 6, 5, 7, 8, 6, 7, 5, 8, 7, 7, 8, 13, 5, 10, 7, 6, 7, 7, 6, 7, 7, 7, 8, 4, 9, 7, 4, 5, 4, 6, 8, 8, 6, 10, 3, 5, 10, 8, 6, 6, 11, 7, 6, 6, 5, 5, 8, 4, 6, 4, 13, 4, 11, 7, 5, 7, 6, 6, 7, 5, 5, 6, 5, 10, 5, 5, 8, 10, 10, 6, 6, 7, 6, 8, 5, 5, 2, 5, 5, 11, 6, 6, 8, 13, 2, 3, 5, 4, 6, 5, 4, 5, 5, 5, 11, 5, 8, 7, 8, 7, 5, 6, 5, 6, 10, 3, 9, 5, 4, 6, 6, 9, 8, 6, 9, 6, 7, 5, 6, 3, 6, 9, 8, 7, 7, 4, 5, 8, 5, 8, 8, 7, 6, 8, 14, 6, 4, 7, 3, 9, 5, 6, 5, 5, 6, 7, 3, 9, 9, 5, 6, 6, 4, 4, 9, 7, 5, 4, 5, 15, 8, 7, 9, 6, 6, 5, 7, 6, 8, 4, 4, 5, 5, 3, 5, 3, 4, 4, 4, 7, 12, 8, 9, 9, 6, 3, 6, 4, 7, 7, 9, 4, 6, 9, 5, 7, 5, 10, 5, 10, 6, 9, 4, 6, 8, 5, 8, 12, 10, 5, 7, 6, 7, 10, 7, 9, 6, 7, 5, 6, 6, 8, 6, 6, 8, 4, 6, 6, 9, 6, 6, 7, 4, 4, 3, 8, 10, 6, 6, 25, 8, 8, 5, 5, 4, 7, 7, 5, 7, 6, 7, 7, 6, 6, 5, 8, 6, 6, 7, 6, 8, 5, 4, 5, 8, 6, 12, 6, 7, 8, 4, 4, 7, 7, 9, 9, 14, 3, 10, 6, 6, 5, 7, 14, 5, 8, 4, 8, 8, 6, 6, 4, 6, 10, 14, 8, 5, 7, 6, 9, 5, 6, 7, 7, 5, 7, 5, 6, 9, 6, 6, 8, 7, 3, 6, 5, 9, 4, 4, 6, 13, 4, 6, 4, 5, 5, 7, 6, 7, 14, 3, 5, 11, 6, 7, 7, 7, 5, 5, 6, 14, 7, 7, 7, 5, 3, 4, 8, 4, 6, 8, 2, 6, 10, 5, 12, 8, 9, 6, 5, 13, 6, 8, 5, 2, 5, 1, 5, 6, 5, 5, 4, 9, 6, 7, 3, 8, 5, 6, 7, 6, 7, 8, 7, 3, 8, 6, 7, 5, 7, 7, 6, 5, 7, 11, 9, 6, 6, 3, 4, 9, 8, 8, 8, 6, 5, 6, 5, 7, 15, 8, 10, 9, 10, 6, 5, 7, 6, 10, 6, 5, 12, 5, 5, 8, 8, 9, 4, 7, 4, 4, 8, 15, 6, 4, 12, 6, 6, 4, 6, 6, 8, 4, 7, 8, 6, 6, 8, 4, 5, 9, 7, 6, 6, 7, 7, 6, 6, 7, 9, 6, 7, 6, 8, 5, 5, 5, 10, 8, 6, 6, 7, 5, 6, 6, 6, 8, 8, 7, 6, 8, 5, 6, 7, 7, 7, 4, 7, 6, 6, 4, 11, 4, 7, 6, 5, 9, 10, 8, 6, 7, 6, 7, 6, 6, 7, 4, 7, 7, 5, 7, 7, 6, 9, 6, 6, 6, 7, 8, 4, 5, 3, 7, 5, 6, 8, 6, 6, 6, 16, 6, 5, 15, 10, 6, 7, 9, 7, 7, 8, 5, 9, 6, 5, 5, 4, 9, 11, 6, 6, 6, 8, 5, 5, 6, 4, 5, 7, 8, 8, 5, 4, 3, 8, 5, 4, 4, 5, 5, 6, 5, 3, 10, 8, 5, 9, 9, 6, 5, 3, 5, 6, 7, 7, 8, 5, 8, 6, 6, 4, 5, 4, 5, 9, 9, 4, 7, 5, 6, 9, 7, 4, 8, 8, 7, 6, 10, 7, 8, 11, 5, 7, 5, 6, 7, 9, 8, 7, 7, 8, 10, 3, 4, 6, 7, 7, 5, 7, 6, 5, 6, 9, 10, 3, 7, 5, 7, 8, 9, 5, 6, 2, 9, 7, 7, 4, 6, 6, 9, 7, 9, 6, 6, 7, 6, 7, 6, 5, 7, 8, 8, 5, 6, 6, 8, 6, 6, 6, 8, 7, 8, 6, 6, 8, 5, 7, 8, 4, 6, 5, 7, 8, 7, 7, 9, 5, 1, 8, 7, 5, 5, 7, 8, 5, 8, 6, 6, 4, 6, 6, 6, 7, 5, 5, 4, 7, 7, 7, 7, 6, 7, 5, 6, 6, 7, 6, 5, 6, 7, 9, 6, 7, 7, 4, 7, 7, 5, 6, 8, 5, 7, 7, 7, 7, 6, 7, 6, 14, 6, 9, 9, 10, 5, 6, 7, 7, 9, 4, 8, 10, 4, 8, 6, 5, 5, 6, 6, 6, 7, 7, 6, 3, 5, 10, 8, 9, 6, 8, 1, 4, 6, 7, 5, 7, 8, 7, 7, 7, 6, 5, 7, 5, 4, 10, 7, 9, 5, 8, 10, 7, 5, 9, 6, 6, 7, 8, 7, 7, 6, 7, 5, 6, 5, 8, 9, 6, 6, 6, 10, 5, 6, 7, 6, 8, 6, 7, 7, 6, 9, 5, 8, 4, 5, 7, 6, 5, 4, 4, 7, 8, 5, 5, 7, 5, 8, 7, 9, 6, 10, 6, 8, 8, 6, 7, 6, 6, 7, 9, 6, 5, 5, 8, 6, 7, 10, 6, 4, 14, 3, 6, 6, 8, 8, 7, 9, 6, 6, 10, 6, 8, 7, 6, 8, 6, 4, 5, 5, 5, 8, 4, 4, 6, 4, 6, 11, 4, 5, 6, 4, 6, 7, 5, 5, 3, 5, 6, 6, 9, 8, 9, 6, 6, 6, 6, 6, 8, 7, 7, 6, 4, 6, 5, 8, 6, 6, 5, 7, 8, 5, 7, 7, 6, 7, 3, 7, 4, 6, 5, 6, 5, 6, 6, 7, 10, 6, 6, 9, 7, 14, 6, 5, 7, 9, 8, 8, 9, 5, 4, 9, 4, 6, 5, 6, 8, 7, 6, 5, 5, 6, 5, 6, 7, 6, 6, 7, 7, 7, 6, 7, 6, 7, 6, 11, 6, 7, 7, 5, 4, 7, 6, 9, 7, 3, 6, 8, 5, 6, 3, 6, 7, 6, 3, 7, 11, 5, 7, 6, 6, 5, 6, 5, 7, 6, 8, 8, 9, 6, 6, 6, 7, 7, 10, 7, 9, 7, 6, 9, 8, 7, 6, 6, 5, 7, 8, 5, 7, 6, 6, 6, 9, 7, 4, 8, 7, 8, 7, 6, 6, 10, 5, 8, 4, 8, 5, 6, 7, 7, 7, 8, 7, 6, 9, 6, 7, 5, 4, 5, 7, 7, 5, 7, 7, 6, 6, 5, 7, 6, 5, 7, 5, 6, 8, 8, 5, 5, 5, 10, 6, 5, 6, 4, 9, 5, 6, 9, 8, 8, 7, 6, 8, 10, 6, 11, 6, 5, 10, 6, 8, 8, 7, 6, 10, 5, 7, 9, 6, 4, 8, 9, 7, 6, 6, 6, 5, 10, 6, 9, 5, 7, 7, 8, 6, 7, 5, 7, 6, 4, 7, 6, 8, 8, 5, 7, 6, 4, 7, 4, 5, 8, 5, 5, 9, 8, 5, 8, 6, 4, 7, 5, 3, 6, 6, 4, 7, 3, 5, 4, 4, 7, 8, 8, 7, 10, 8, 6, 6, 7, 5, 6, 5, 7, 8, 4, 7, 9, 5, 7, 5, 5, 7, 7, 7, 5, 5, 5, 6, 9, 6, 6, 8, 7, 5, 5, 7, 6, 8, 6, 8, 10, 9, 7, 8, 5, 6, 6, 7, 6, 8, 7, 6, 8, 5, 7, 5, 6, 7, 6, 5, 8, 5, 7, 3, 7, 6, 7, 8, 8, 4, 6, 2, 8, 6, 6, 5, 7, 5, 8, 8, 9, 7, 7, 7, 8, 7, 8, 6, 5, 8, 11, 10, 7, 7, 4, 6, 8, 5, 4, 8, 3, 5, 6, 8, 9, 7, 4, 5, 8, 8, 5, 5, 6, 6, 6, 7, 9, 6, 6, 11, 4, 7, 5, 9, 9, 6, 8, 6, 6, 5, 5, 8, 7, 7, 5, 7, 6, 12, 7, 6, 7, 5, 1, 10, 5, 3, 7, 5, 5, 6, 6, 7, 8, 8, 7, 5, 3, 5, 7, 7, 8, 9, 4, 5, 8, 8, 6, 5, 7, 7, 6, 7, 5, 8, 11, 5, 5, 4, 5, 5, 1, 9, 6, 9, 9, 5, 6, 7, 7, 9, 6, 7, 7, 7, 5, 4, 5, 6, 6, 5, 6, 4, 6, 6, 5, 5, 3, 5, 7, 4, 6, 4, 8, 6, 6, 6, 6, 6, 7, 4, 3, 4, 12, 6, 6, 6, 6, 8, 6, 6, 7, 12, 8, 5, 11, 4, 6, 6, 5, 6, 7, 6, 5, 7, 7, 10, 6, 5, 7, 6, 5, 6, 6, 6, 6, 5, 10, 19, 7, 7, 8, 5, 6, 9, 6, 6, 12, 5, 4, 5, 3, 12, 4, 6, 4, 7, 4, 9, 4, 5, 3, 4, 7, 9, 6, 5, 7, 8, 5, 6, 5, 4, 8, 7, 5, 7, 5, 4, 7, 6, 4, 7, 5, 5, 7, 6, 7, 8, 11, 5, 5, 8, 3, 5, 4, 6, 6, 3, 7, 7, 5, 6, 9, 12, 7, 5, 5, 6, 9, 5, 7, 10, 6, 9, 5, 6, 6, 6, 7, 8, 6, 5, 7, 5, 7, 5, 5, 5, 8, 6, 5, 5, 5, 7, 5, 5, 5, 7, 5, 5, 6, 10, 8, 7, 7, 5, 6, 6, 5, 12, 7, 8, 6, 6, 5, 8, 5, 5, 5, 5, 6, 5, 9, 10, 6, 6, 5, 4, 5, 4, 6, 6, 5, 7, 5, 7, 7, 4, 7, 5, 9, 6, 6, 4, 6, 6, 6, 5, 6, 6, 6, 6, 4, 8, 3, 5, 9, 7, 9, 6, 4, 12, 6, 7, 6, 7, 6, 8, 16, 7, 7, 5, 10, 7, 8, 6, 6, 7, 7, 11, 6, 11, 5, 6, 9, 5, 8, 6, 7, 5, 6, 7, 6, 8, 9, 7, 2, 10, 5, 7, 6, 7, 6, 6, 6, 6, 10, 4, 4, 6, 8, 6, 6, 9, 7, 6, 2, 6, 7, 5, 7, 5, 15, 8, 6, 8, 4, 6, 7, 7, 7, 8, 8, 7, 5, 5, 6, 5, 7, 7, 7, 5, 6, 9, 8, 9, 5, 6, 4, 6, 5, 5, 7, 5, 8, 5, 4, 5, 5, 7, 7, 7, 5, 6, 8, 9, 4, 6, 6, 11, 6, 8, 9, 6, 5, 6, 6, 8, 6, 5, 7, 6, 5, 7, 4, 9, 5, 4, 6, 6, 6, 3, 7, 6, 6, 5, 13, 7, 6, 8, 5, 7, 5, 5, 8, 7, 7, 7, 7, 6, 9, 10, 8, 9, 6, 6, 7, 10, 6, 7, 7, 6, 6, 6, 6, 5, 7, 7, 5, 6, 4, 6, 7, 9, 8, 4, 9, 5, 5, 6, 4, 7, 8, 9, 7, 2, 4, 7, 7, 7, 7, 7, 11, 8, 9, 6, 4, 6, 6, 5, 8, 7, 7, 6, 7, 8, 8, 8, 6, 6, 8, 9, 8, 9, 7, 7, 5, 7, 5, 9, 6, 4, 6, 7, 4, 9, 7, 4, 4, 5, 9, 10, 8, 10, 9, 5, 6, 6, 5, 7, 7, 6, 5, 6, 9, 8, 7, 4, 6, 6, 7, 4, 5, 7, 7, 5, 9, 6, 7, 6, 9, 9, 8, 7, 8, 5, 3, 4, 11, 14, 5, 5, 4, 3, 6, 6, 5, 6, 7, 6, 8, 8, 9, 7, 6, 2, 6, 6, 5, 5, 5, 3, 8, 8, 11, 7, 6, 7, 6, 6, 7, 8, 6, 7, 7, 6, 6, 8, 8, 5, 5, 16, 8, 6, 7, 9, 5, 8, 6, 5, 4, 7, 5, 6, 8, 7, 5, 6, 7, 6, 6, 7, 10, 7, 10, 9, 8, 7, 5, 8, 5, 8, 7, 7, 9, 9, 5, 7, 6, 6, 5, 7, 5, 6, 8, 7, 6, 6, 10, 6, 5, 5, 8, 5, 7, 5, 5, 7, 6, 6, 5, 6, 0, 6, 11, 5, 5, 9, 6, 5, 9, 5, 6, 6, 6, 6, 6, 7, 7, 11, 10, 4, 7, 5, 8, 8, 8, 5, 6, 6, 7, 8, 7, 8, 5, 5, 5, 6, 7, 7, 5, 8, 5, 5, 5, 10, 8, 5, 10, 5, 4, 5, 5, 6, 5, 9, 5, 4, 5, 7, 4, 4, 6, 15, 7, 6, 8, 7, 10, 7, 7, 5, 6, 7, 6, 6, 6, 8, 5, 6, 4, 2, 6, 5, 8, 8, 5, 6, 5, 7, 10, 7, 8, 7, 6, 8, 7, 11, 7, 6, 6, 8, 7, 10, 16, 6, 7, 6, 4, 5, 4, 4, 7, 8, 8, 5, 6, 4, 6, 3, 5, 9, 8, 4, 6, 4, 6, 3, 6, 9, 7, 8, 3, 14, 6, 8, 4, 7, 6, 5, 6, 4, 8, 5, 7, 7, 7, 7, 5, 5, 7, 5, 6, 6, 7, 7, 7, 7, 4, 11, 6, 7, 4, 3, 13, 5, 8, 9, 8, 6, 8, 7, 7, 6, 7, 8, 6, 6, 9, 5, 8, 6, 7, 11, 6, 5, 8, 7, 9, 5, 7, 5, 7, 8, 4, 4, 8, 5, 7, 5, 5, 6, 7, 9, 7, 6, 11, 8, 3, 5, 7, 11, 4, 6, 7, 7, 7, 6, 5, 5, 7, 5, 6, 6, 7, 6, 5, 7, 8, 6, 5, 7, 14, 10, 6, 7, 7, 5, 6, 11, 5, 6, 5, 10, 4, 3, 6, 5, 8, 6, 6, 6, 7, 7, 8, 5, 7, 6, 5, 8, 7, 9, 6, 5, 6, 8, 5, 7, 7, 7, 6, 7, 6, 8, 5, 7, 9, 8, 5, 7, 5, 8, 8, 9, 12, 7, 4, 7, 10, 8, 7, 7, 6, 6, 7, 6, 7])Rook Weights
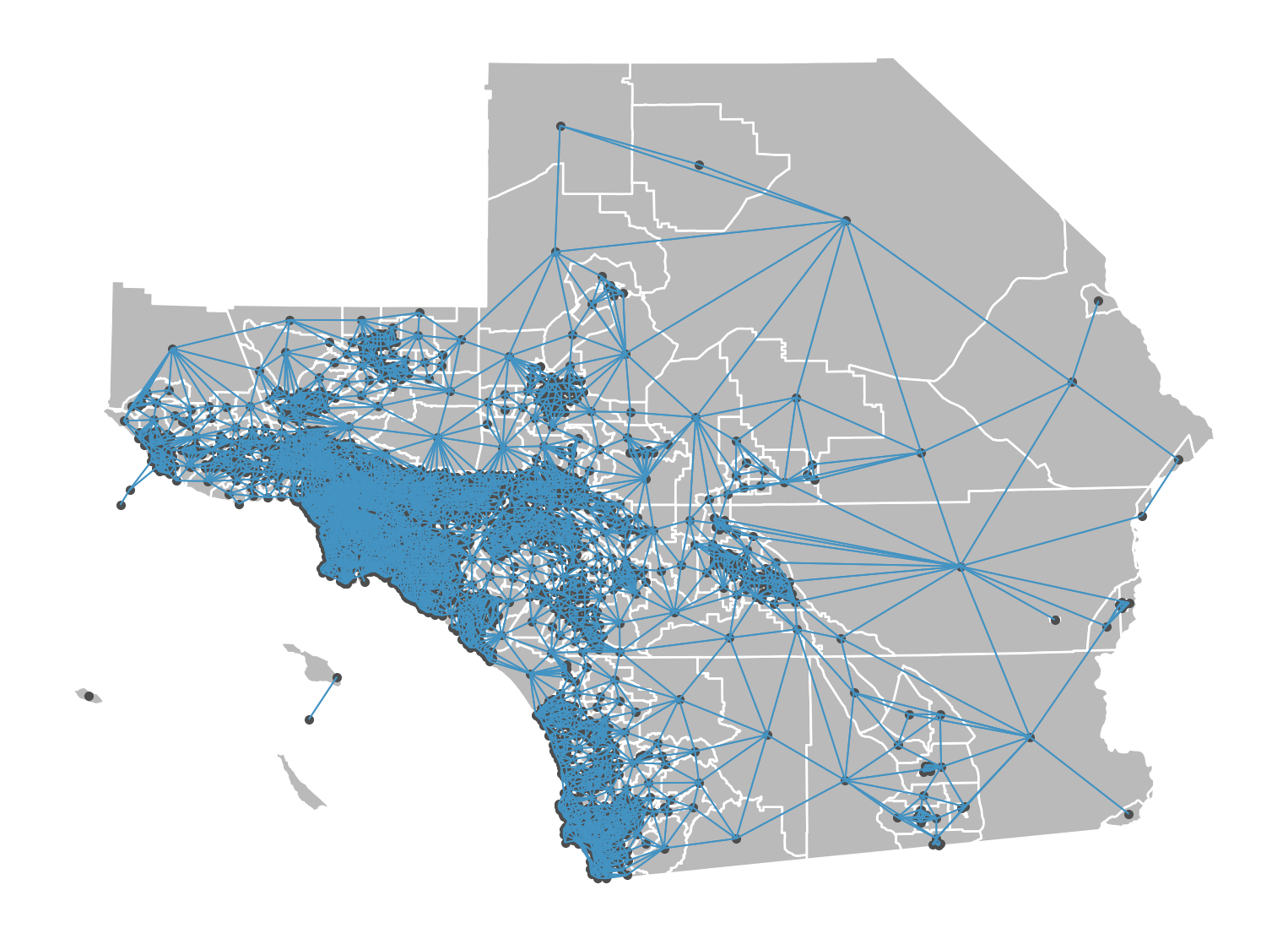
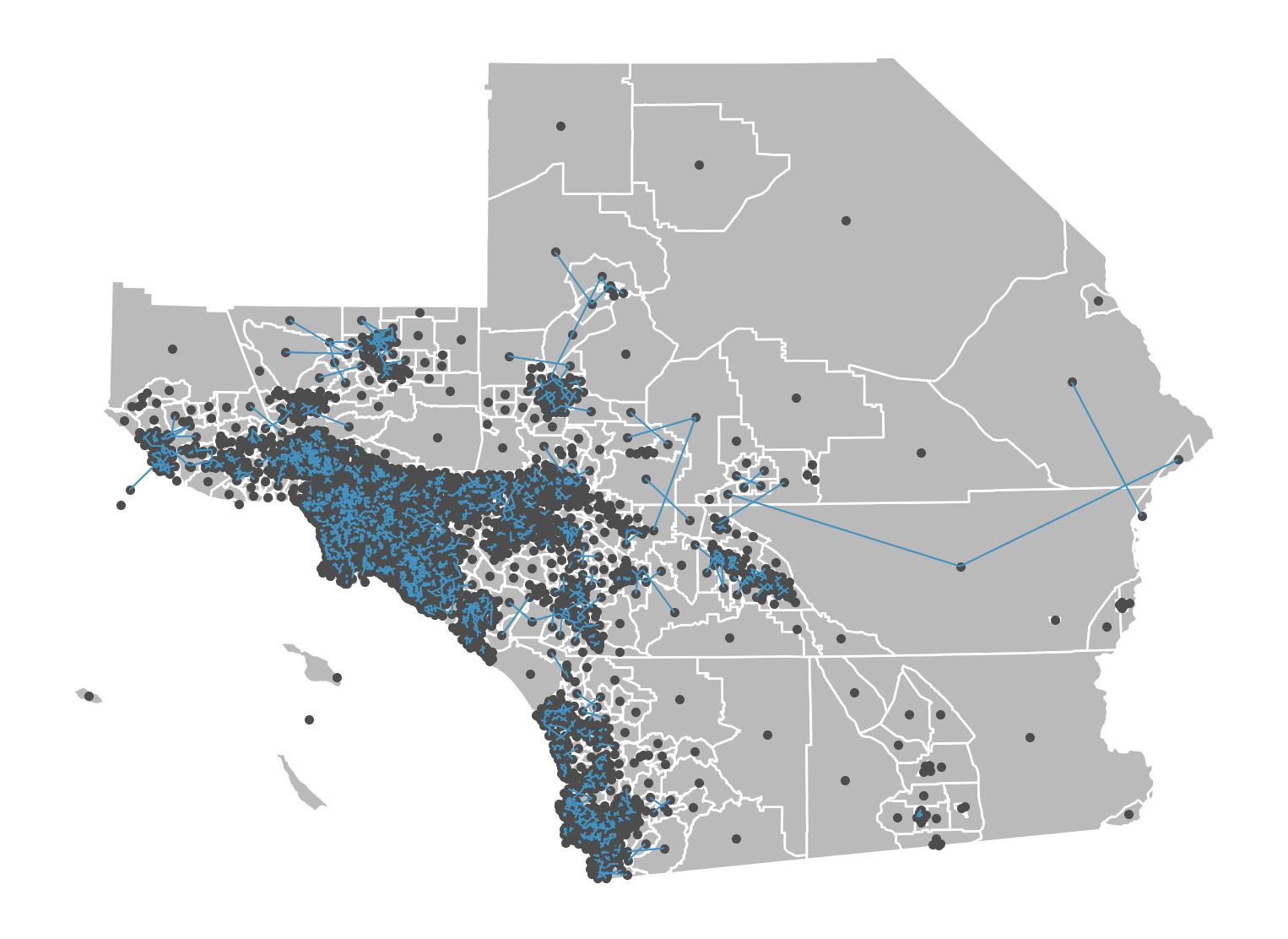
Rook weights are another type of contiguity weight, but consider observations as neighboring only when they share an edge. The rook neighbors of an observation may be different than its queen neighbors, depending on how the observation and its nearby polygons are configured.
We can construct this in the same way as the queen weights:
Rook Graph
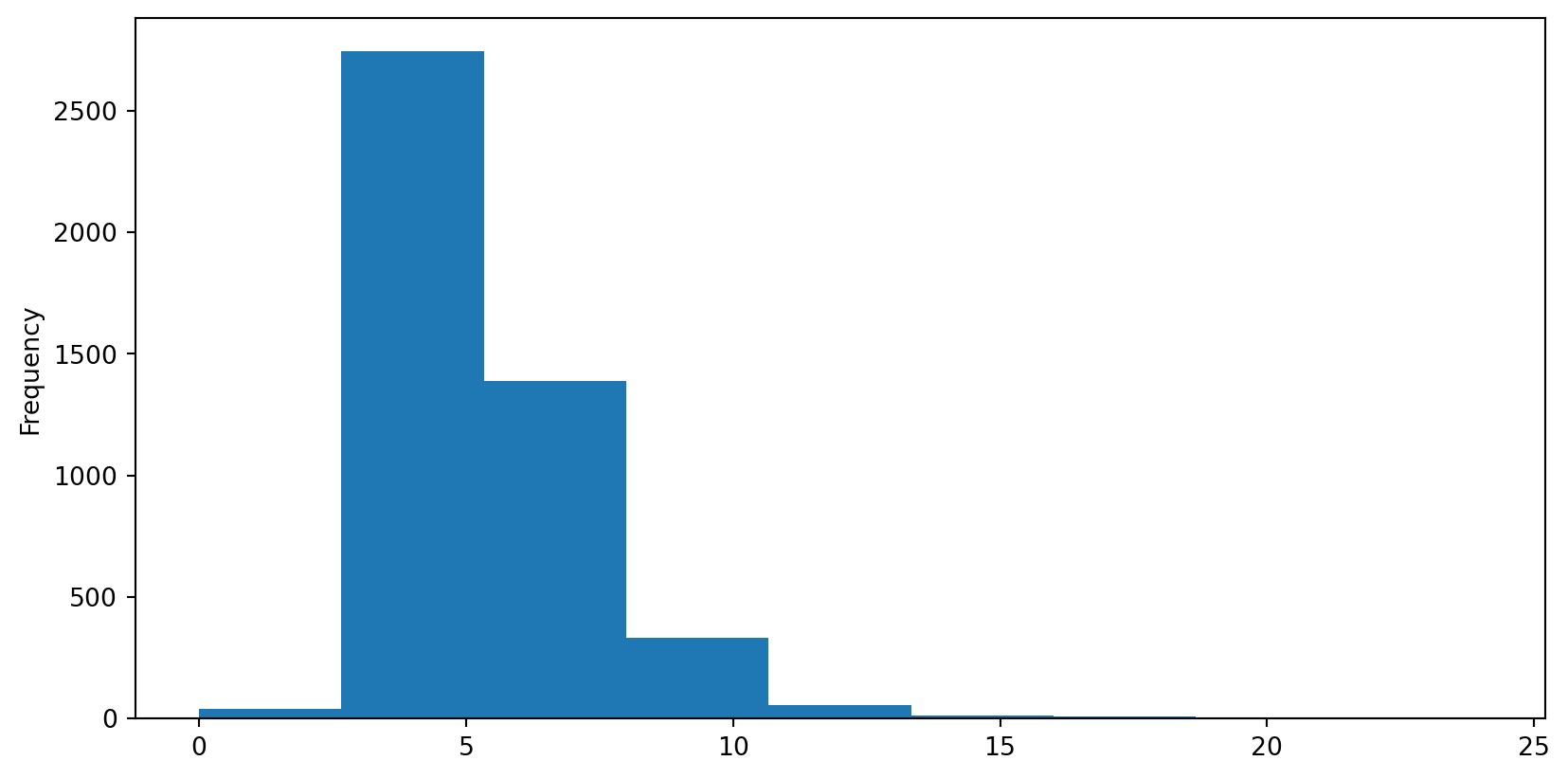
Connectivity Histogram
Bishop Weights
In theory, a “Bishop” weighting scheme is one that arises when only polygons that share vertexes are considered to be neighboring. But, since Queen contiguigy requires either an edge or a vertex and Rook contiguity requires only shared edges, the following relationship is true:
\[ \mathcal{Q} = \mathcal{R} \cup \mathcal{B} \]
where \(\mathcal{Q}\) is the set of neighbor pairs via queen contiguity, \(\mathcal{R}\) is the set of neighbor pairs via Rook contiguity, and \(\mathcal{B}\) via Bishop contiguity.
Bishop Weights
\[ \mathcal{Q} \setminus \mathcal{R} = \mathcal{B}\]
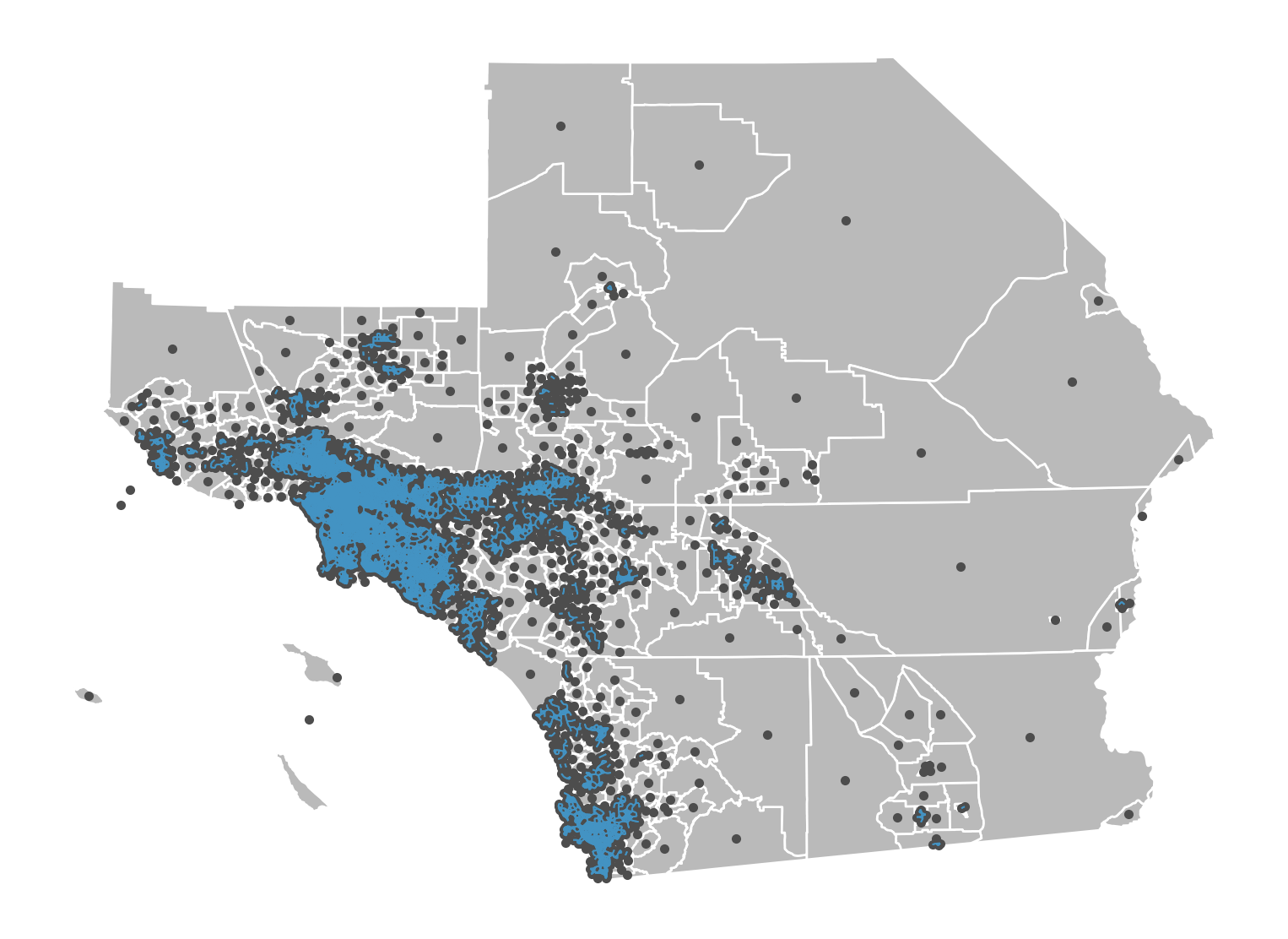
Bishop weights entail all Queen neighbor pairs that are not also Rook neighbors.
Bishop Weights
PySAL does not have a dedicated bishop weights constructor, but you can construct very easily using the w_difference function. This function is one of a family of tools to work with weights, all defined in libpysal.weights, that conduct these types of set operations between weight objects.
Bishop Graph
Connectivity
[(np.int64(0), np.int64(1624)),
(np.int64(1), np.int64(1728)),
(np.int64(2), np.int64(881)),
(np.int64(3), np.int64(292)),
(np.int64(4), np.int64(55))]Thus, many tracts have no bishop neighbors. But, a few do.
Bishop Connectivity Graph
Distance Based Weights
Distance
There are many other kinds of weighting functions in PySAL. Another separate type use a continuous measure of distance to define neighborhoods.
<Projected CRS: EPSG:26911>
Name: NAD83 / UTM zone 11N
Axis Info [cartesian]:
- E[east]: Easting (metre)
- N[north]: Northing (metre)
Area of Use:
- name: North America - between 120°W and 114°W - onshore and offshore. Canada - Alberta; British Columbia; Northwest Territories; Nunavut. United States (USA) - California; Idaho; Nevada, Oregon; Washington.
- bounds: (-120.0, 30.88, -114.0, 83.5)
Coordinate Operation:
- name: UTM zone 11N
- method: Transverse Mercator
Datum: North American Datum 1983
- Ellipsoid: GRS 1980
- Prime Meridian: GreenwichThreshold Based Neighbors
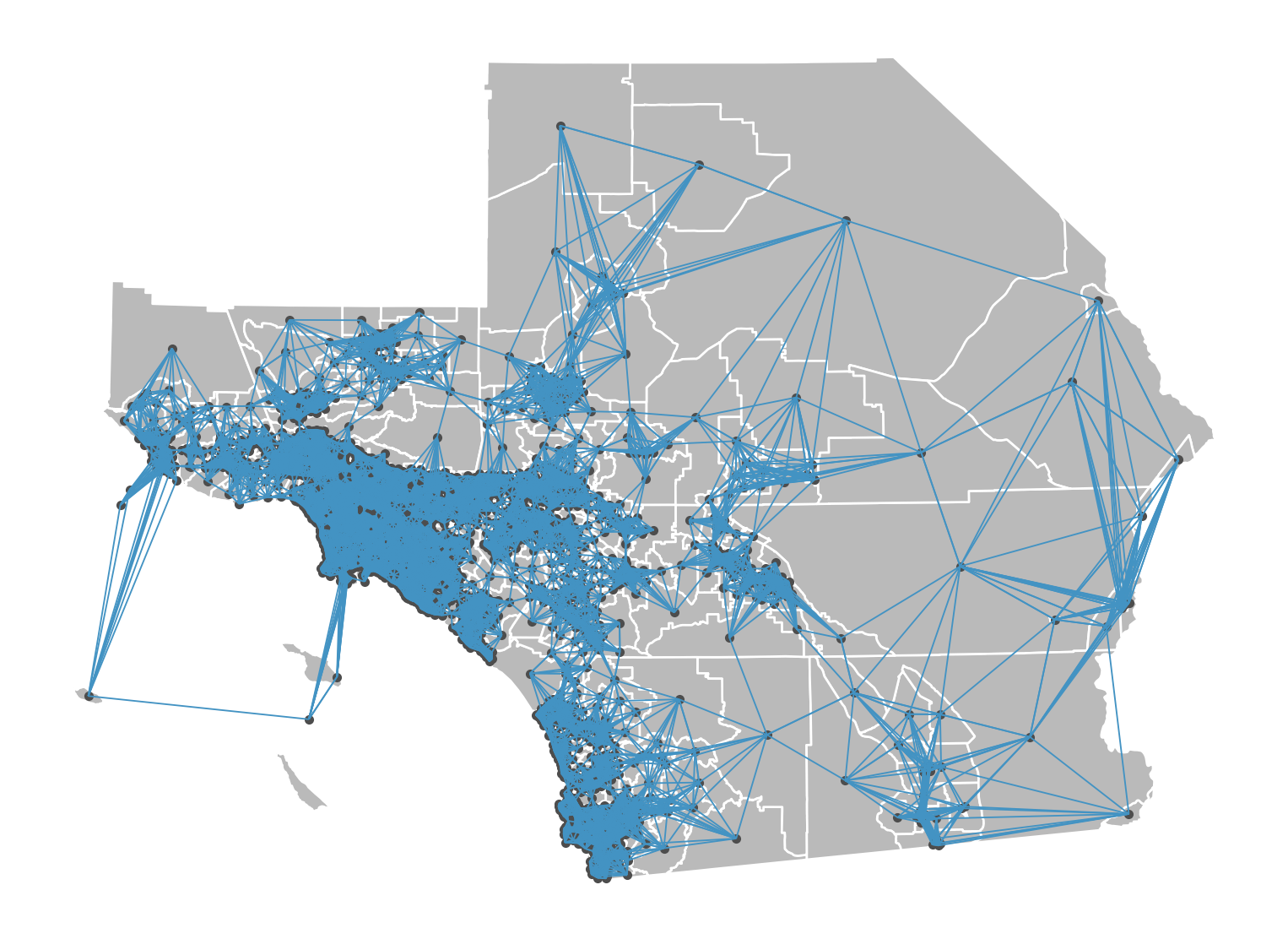
Our coordinate system (UTM 11N) measures distance in meters, so that’s how we’ll define our neighbors
Plotting Threshold Neighbor Graph
knn defined weights
Wrong Way to define KNN
Bad KNN
Correct Way to define KNN
Correct KNN
Comparison
[np.int64(501),
np.int64(2296),
np.int64(2960),
np.int64(974),
np.int64(167),
np.int64(4496),
np.int64(2295),
np.int64(4422)][np.int64(501),
np.int64(2960),
np.int64(2296),
np.int64(974),
np.int64(167),
np.int64(4496),
np.int64(2881),
np.int64(2297)]Kernel W
Kernel Weights are continuous distance-based weights that use kernel densities to define the neighbor relationship. Typically, they estimate a bandwidth, which is a parameter governing how far out observations should be considered neighboring. Then, using this bandwidth, they evaluate a continuous kernel function to provide a weight between 0 and 1.
Many different choices of kernel functions are supported, and bandwidths can either be fixed (constant over all units) or adaptive in function of unit density.
For example, if we want to use adaptive bandwidths for the map and weight according to a gaussian kernel:
Adaptive gaussian kernel weights
bandwidth = the distance to the kth nearest neighbor for each observation
bandwith is changing across observations
Kernel Weights
Kernel Weights (Continuous)
Adaptive bandwidths
Adaptive bandwidths
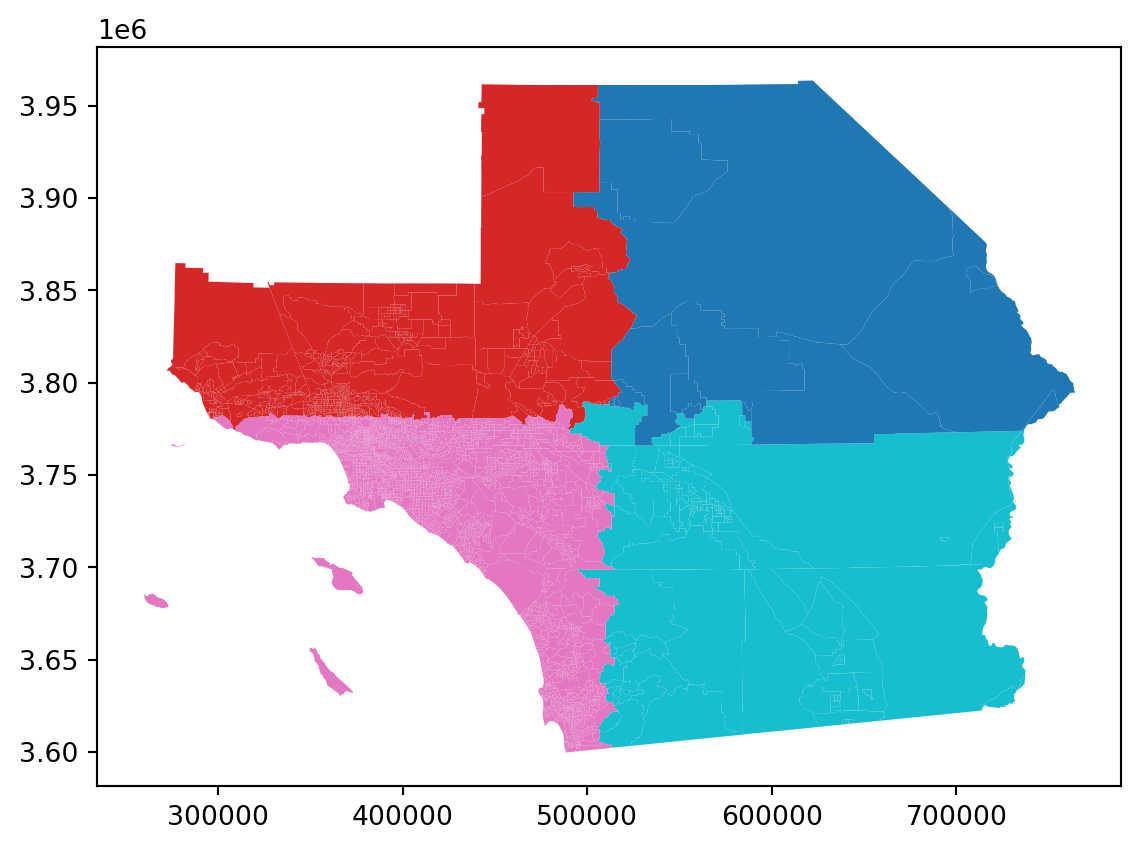
Block Weights
Regions
Regions
Block Weights
Connectivity
Region Membership
| geoid | n_asian_under_15 | n_black_under_15 | n_hispanic_under_15 | n_native_under_15 | n_white_under_15 | n_persons_under_18 | n_asian_over_60 | n_black_over_60 | n_hispanic_over_60 | ... | year | n_total_housing_units_sample | p_nonhisp_white_persons | p_white_over_60 | p_black_over_60 | p_hispanic_over_60 | p_native_over_60 | p_asian_over_60 | p_disabled | geometry | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| quad | |||||||||||||||||||||
| 1 | 15 | 15 | 15 | 15 | 15 | 15 | 15 | 0 | 0 | 0 | ... | 15 | 15 | 15 | 0 | 0 | 0 | 0 | 0 | 0 | 15 |
| 2 | 761 | 761 | 761 | 761 | 761 | 761 | 761 | 0 | 0 | 0 | ... | 761 | 761 | 755 | 0 | 0 | 0 | 0 | 0 | 0 | 761 |
| 3 | 3625 | 3625 | 3625 | 3625 | 3625 | 3625 | 3625 | 0 | 0 | 0 | ... | 3625 | 3625 | 3612 | 0 | 0 | 0 | 0 | 0 | 0 | 3625 |
| 4 | 179 | 179 | 179 | 179 | 179 | 179 | 179 | 0 | 0 | 0 | ... | 179 | 179 | 179 | 0 | 0 | 0 | 0 | 0 | 0 | 179 |
4 rows × 194 columns
Conclusion
Recap of Key Points
- Spatial Weights
- Contiguity Weights
- Distance Based Weights
Questions